Microtubules (MTs), or the "bones" of the cell, give the cell its shape and allow it move and divide. How cells “read and write” unique microtubule populations, or decode the “tubulin code”, remains mysterious.
Cells can use post-translational modifications and microtubule associated proteins, or MAPs, to transform the surface of the microtubule, and, in turn, its identity and function within the cell.
MTs are composed of αβ-tubulin heterodimers that self-assemble. These dimers stack head-to-tail to form protofilaments, with 11-16 protofilaments (PFs) associating laterally to form hollow tubes. In vivo, MTs are largely 13-PF MTs because they are nucleated from the γ-tubulin ring complex, a proposed template for 13-PF MTs, or are highly regulated by MAPs that promote 13-PF MTs, such as end-binding proteins (EBs) and doublecortin which lock PFs in specific arrangements, as seen in vitro.
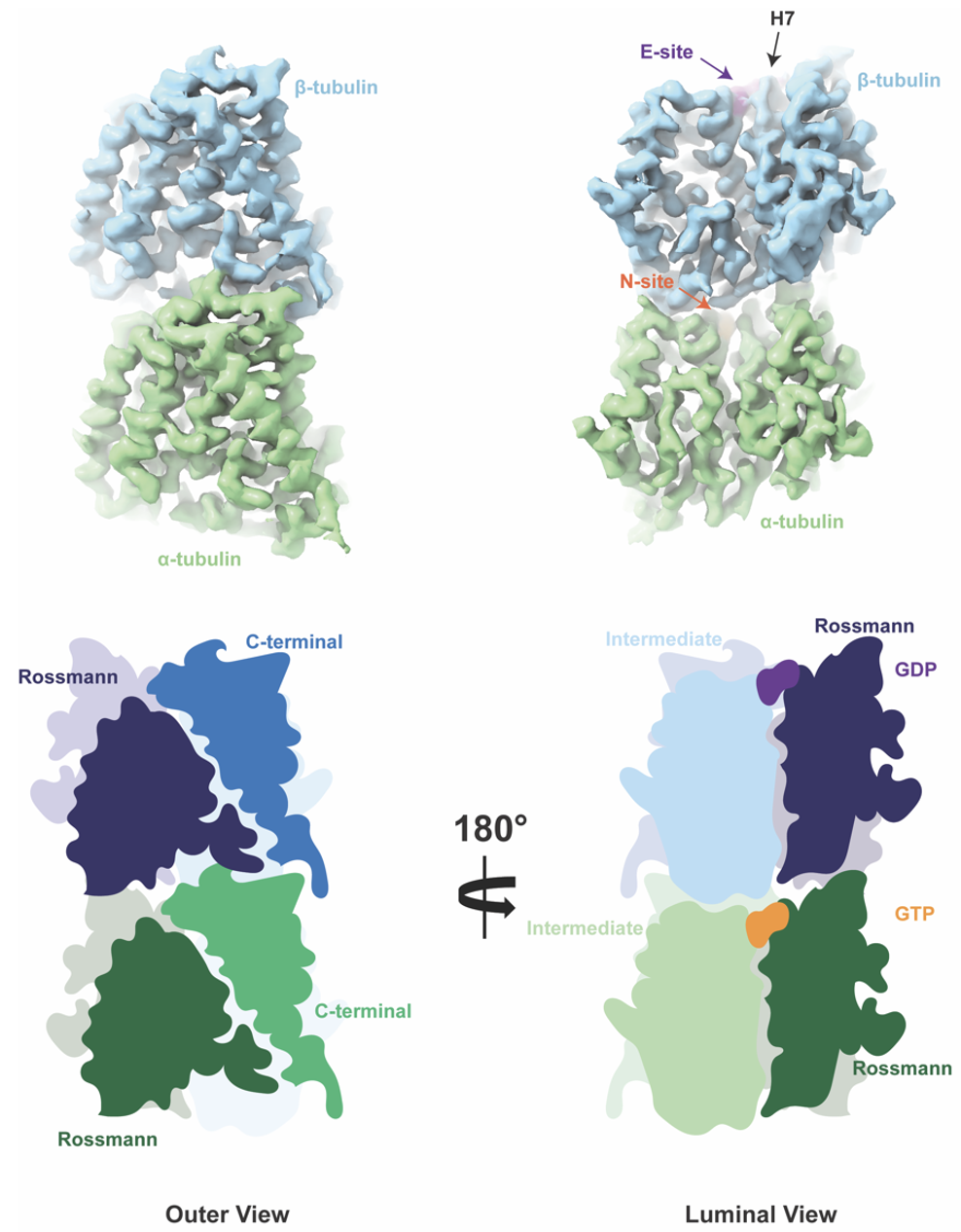
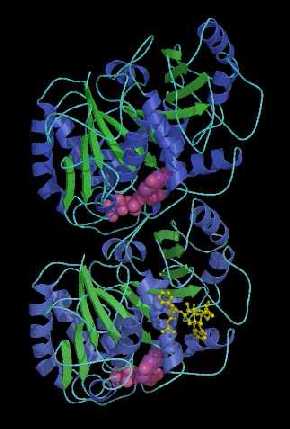
The process of MT assembly is driven by GTP hydrolysis. Thanks to α-tubulin’s built-in GTPase activity, or ability to hydrolyze GTP, MTs can harvest hydrolysis energy and release it explosively to do work. Tubulin belongs to a unique class of GTPases, composed of tubulin, which is 50 kDa found in all eukaryotes, and filamenting temperature sensitive strain Z (FtsZ), which is 40 kDa found in eubacteria and archaea. Tubulin and FtsZ share a common fold of two domains connected by a central helix (H7): the Rossman domain (purple) and the smaller Intermediate domain where the stabilizing chemotherapy agent taxol binds (blue). I spent most of my PhD staring at these domains and wondering what chemical moieties or other proteins stabilize their structure, just like a damper can stabilize a bridge and an abutment, or the component of the bridge that take on pressure and dissipates it onto the Earth.
For my first project, I was intrigued by diseases that exploited these mechanisms. In particular, cancer cells were shown to grow "micro-tentacles", which were highly acetylated microtubules that were so stable and stress-resistthat they could extend like hairs outside of the cell and help tumors grow and proliferate. These "hairs" made the hairs on the back of my neck stand up.
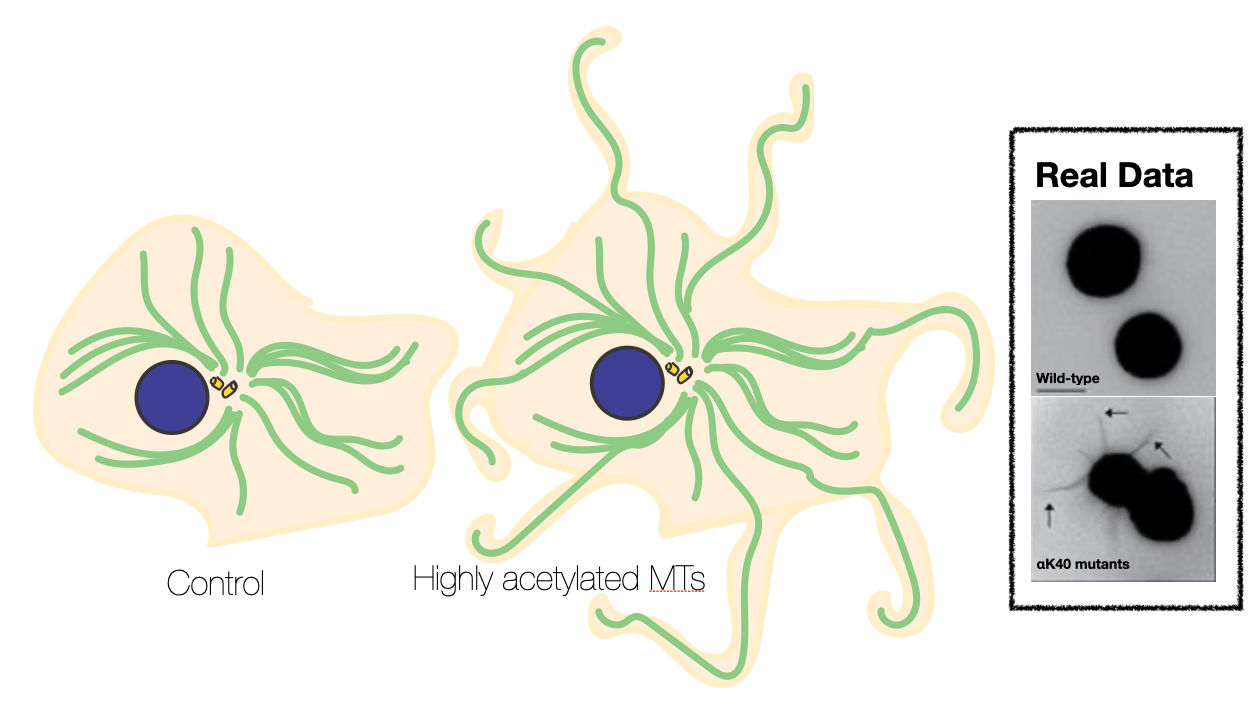
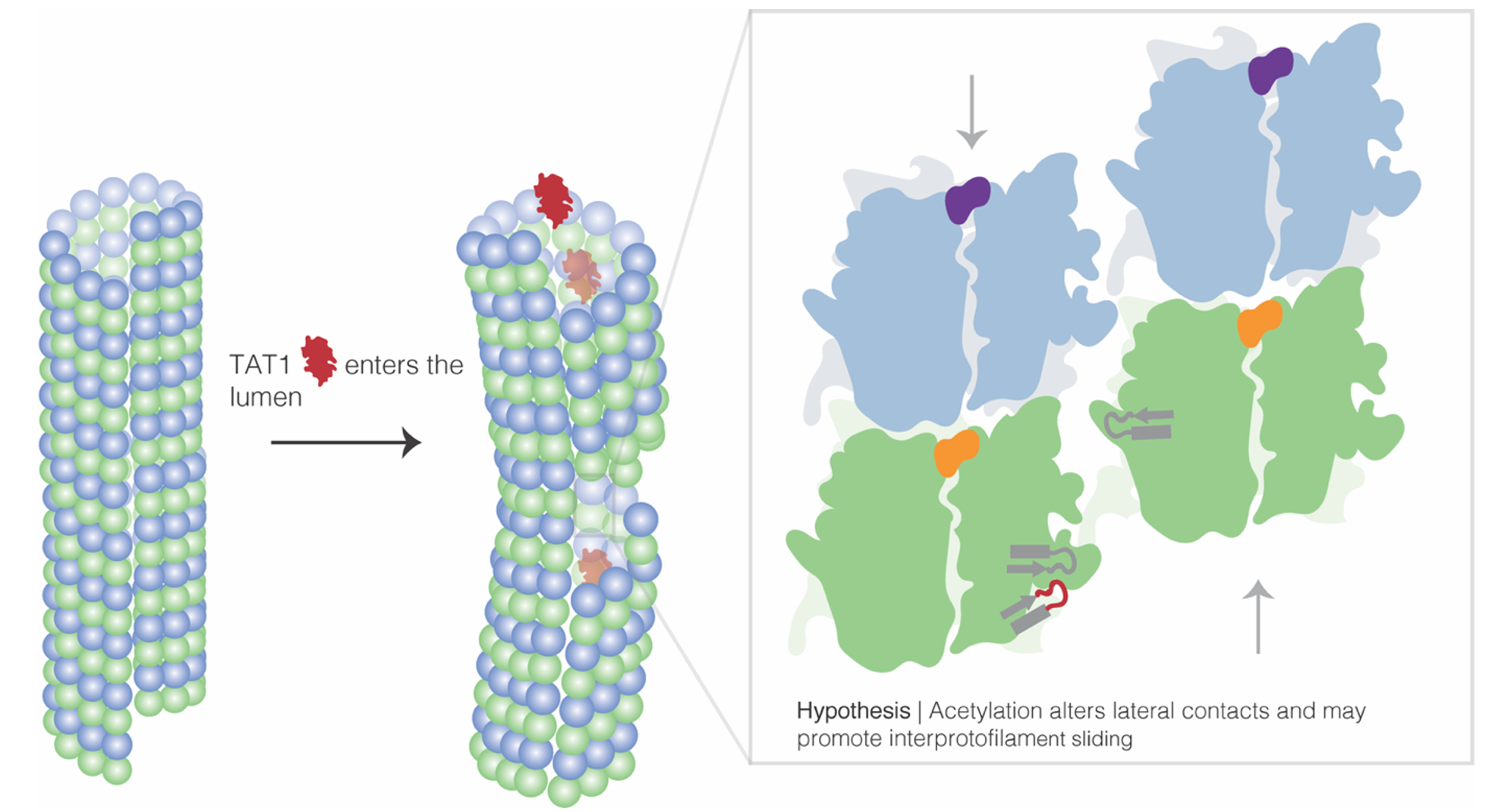
Misregulation of αK40 acetylation has been linked to axonal transport defects associated with Huntington’s disease, Charcot–Marie–Tooth disease, amyotrophic lateral sclerosis, and Parkinson’s disease. However, it was unclear whether these effects were causative or correlative, and if elevated αK40 acetylation levels were a potential biomarker for disease. Using cryo-electron microscopy and molecular dynamics, I provided a mechanistic, structural explanation for how acetylation increases MT stability. I emphasized that by weakening lateral contacts and increasing MT flexibility, acetylation increases inter-protofilament sliding, or how neighboring protofilaments slide in opposite directions to accommodate mechanical stress, or in other words, aceylation enhanced microtubule tensile strength.
When thinking about this, I was inspired by Inka engineers. Impressively, the Inka leveraged the engineering concept of tensile strength long before the Europeans did. For example, they carefully thought about what materials could be pulled and stretched in opposite directions long enough to support foot traffic across the bridge without tearing or snapping. They explored grass cables as thick as a man's thigh and braided and twisted it in ways that could hold 5,175 pounds, or the combined weight of 12 llamas! Acetylation was a PTM the cell could employ to help microtubules bundle for over a meter long within the body (like in the sciatic nerve!) or extend outward from the cell to create those crazy, hair-like protrusions.
Finally, I resolved and modeled the final part of the tubulin structure, the αK40 loop, which was challenging to visualize due to its flexibility. The majority of the structure was resolved by my advisor Eva Nogales, and her colleagues Sharon Wolf and Kenneth Downing, in the 1990's.
After this project grew it sea legs in 2018, I got excited about multiple PTMs and MAPs working together to brilliantly create unique populations of microtubules, from the thick microtubule bundles that line brain cells to those that form motile cilia and help position eggs within my ovaries - wild! Also, Aim 2 of my qualifying exam! I had big dreams as a 2nd year.
I attended the 2018 EMBO/EMBL Conference titled, "Microtubules: from atoms to complex systems" where I drank German beer the entire day and I saw a beautiful talk by Kassandra Ori-McKenney that introduced me to MAP7, or the MAP to my heart.
Like PTMs, microtubule-associated proteins, or MAPs, are architects of specialized MT populations for diverse functions in the cell. Based on their mode of action, MAPs can be grouped into the following groups: (1) motile MAPs, or motor proteins, such as kinesin and dynein, that generate forces and movement; (2) enzymes that break or depolymerize MTs; (3) MT nucleators (4) end-binding proteins that specifically associate with +/- ends of MTs; and (5) “structural” MAPs. Structural MAPs can directly shape the cytoskeleton in ways such as promoting MT bundling, cross-linking multiple cytoskeletal filaments, or stabilizing and polymerizing MTs.
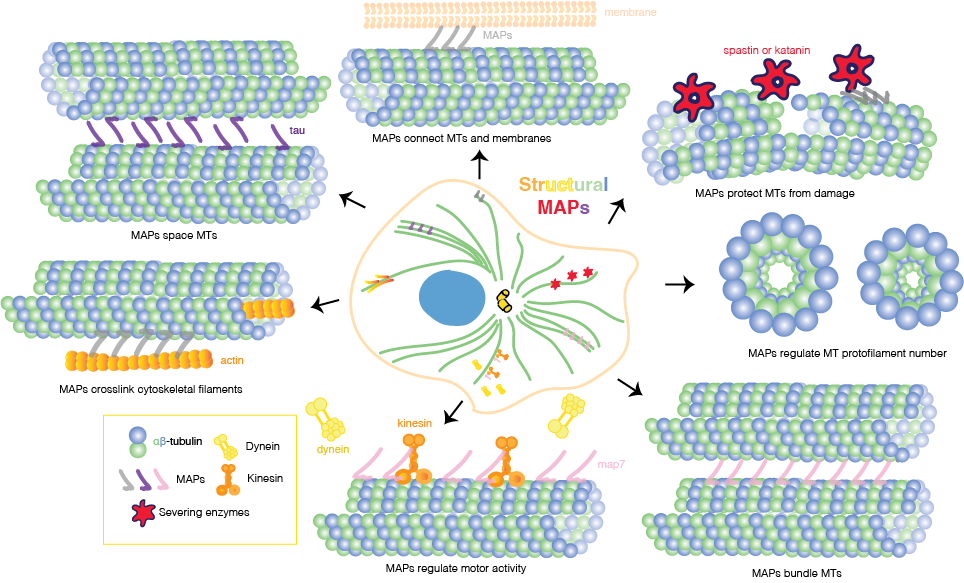
They share a positively charged MT-binding domain (MTBD) and projection domain and not only influence MT structure and behavior but may also drive interactions with other MAPs. Since their discovery in the 1970s, multiple hypotheses have formed to describe how structural MAPs can anchor and manipulate MT populations. First, MAPs could physically link neighboring MTs or, as a second hypothesis, MAP stabilization of MTs could be enough to trigger bundling, for instance, by shielding the negatively charged C-terminal tails, reducing electrostatic repulsion between MTs, and therefore promoting spontaneous MT bundling. When the C-terminal tails are cleaved from the MT surface using subtilisin, thus neutralizing the negative surface charge along the MT lattice, scientists see that MTs bundle immediately, which provides strong evidence for the latter hypothesis.
MAP7 was a structural MAP, that together with cryo-EM and single molecule biophysics, my amazing team and I could resolve to high resolution and catch in action.
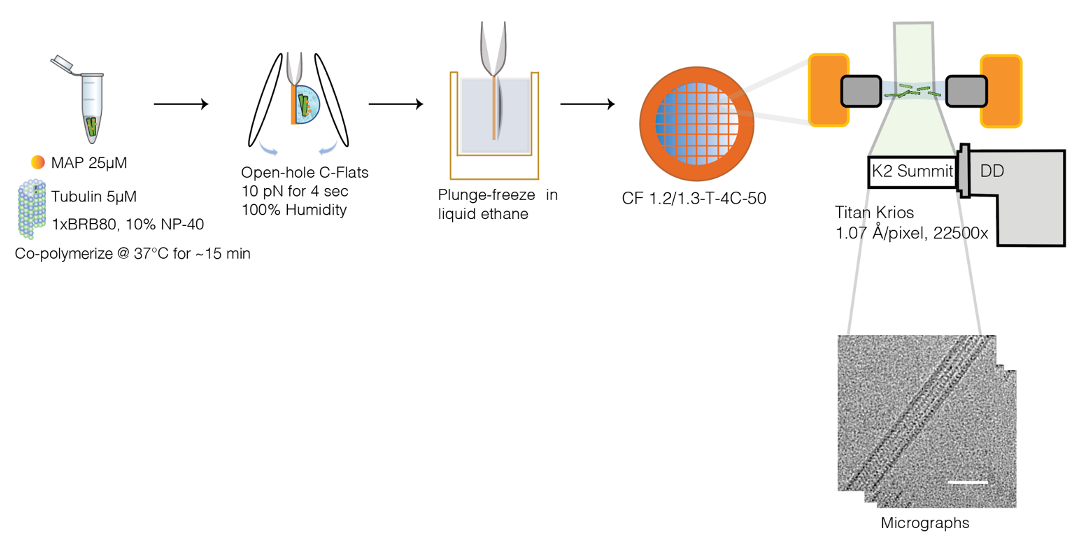
Intriguingly, how MAPs interact with each other to modulate MT function and how this interplay causes disease remains elusive. Alzheimer’s can develop when tau, one of the cell’s most abundant MAPs, is misregulated and, instead, creates filamentous aggregates called neurofibrillary tangles. Notably, MAP7 was shown to kick tau off the MT surface, but recruit the motor kinesin—harboring this elusive, dual molecular role.
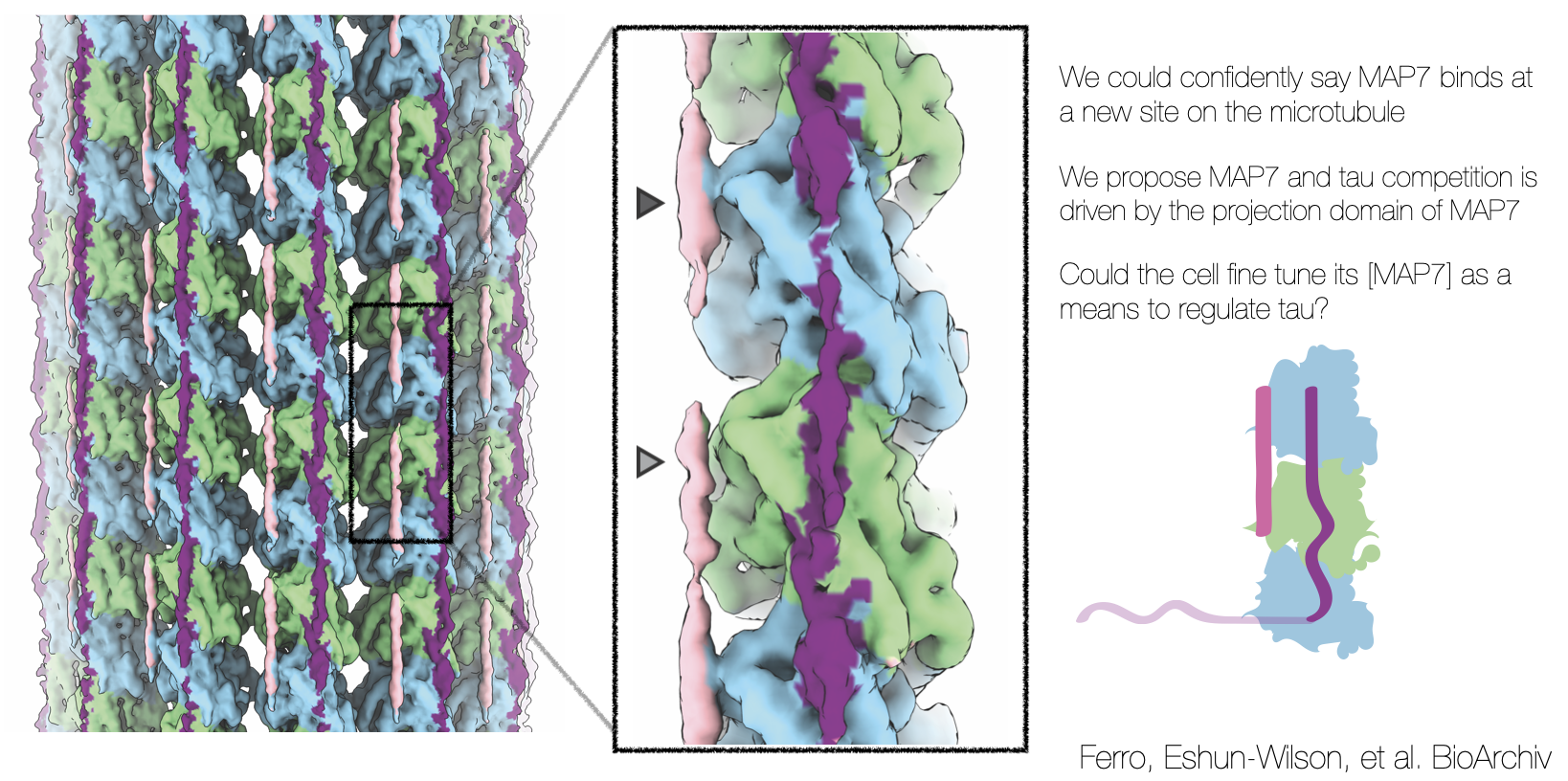
After, talking about this project idea to all my friends, I discovered that my friend Luke Ferro was also interested in MAP7 and had already purified it! Along with tau! How perfect! Together, we started planning experiments. How did molecular motors like kinesin walk along a MAP7-decorated microtubule? Or a MAP7-decorated and highly acetylated microtubule? What about MAP7 and tau? Could they co-bind?
I observed that while MAP7 and tau bound the MT in completely different spots, but in the same spot as kinesin and dynein. It was almost like a game of legos, who binds where and when? How is this MAP competition exploited in disease.
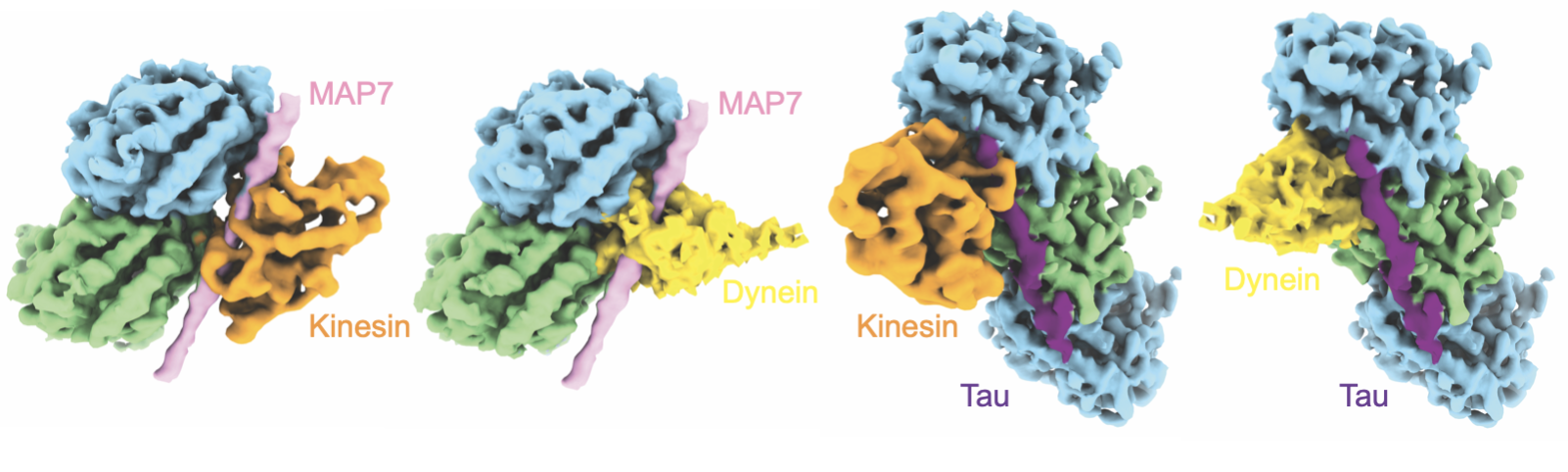
Over time, our team combined more advanced cryo-EM image processing techniques (the quintessential Qianglin) and molecular dynamics (the meticulous Mert) to quantitatively improve the quality of the Coulomb potential map and build in the atomic map (the fabulous Frank).
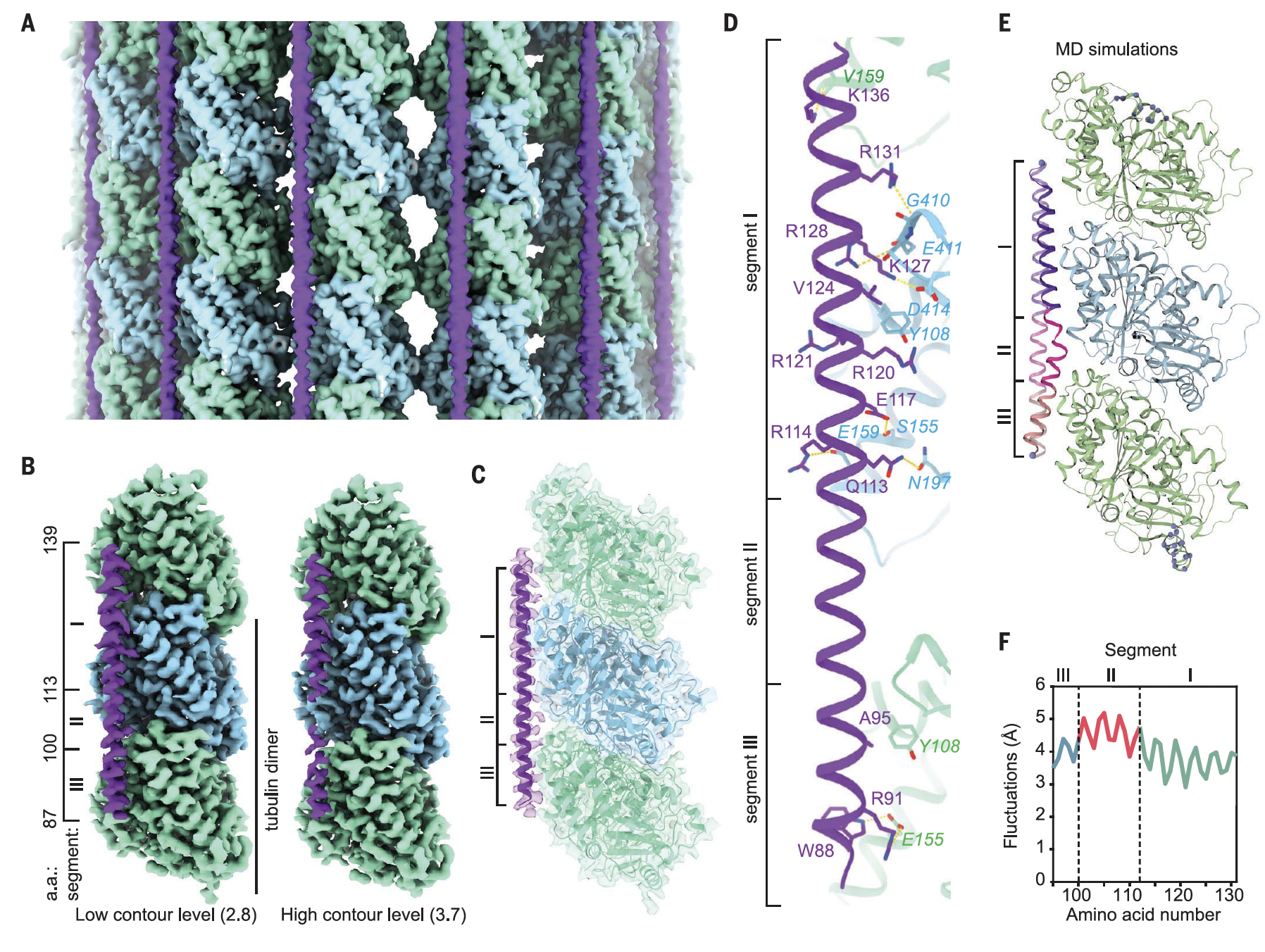
We concluded that MAP7 tethered kinesin via its kinesin-binding domain close enough to the MT lattice, so that it could walk along free sites, when there were free sites available.
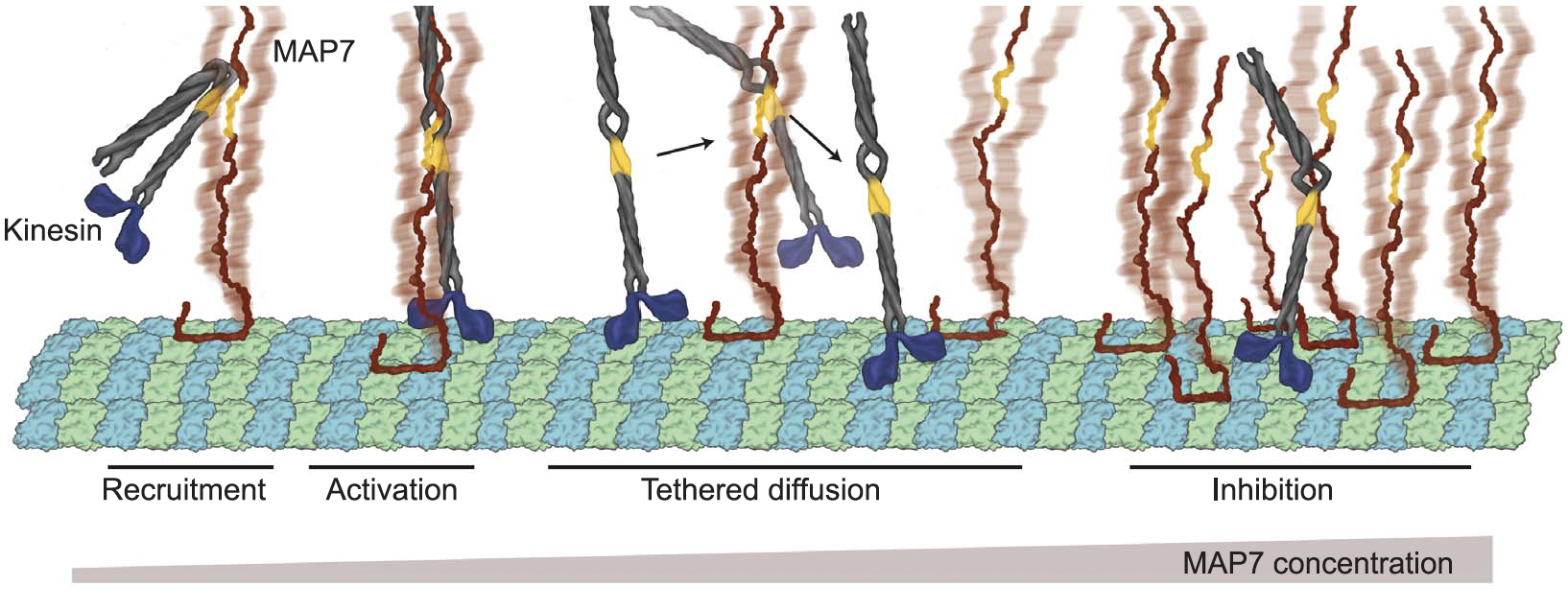
MAP7 was truly the MAP to my heart and open my eyes to the open questions and intriguing complexities of microtubule regulation yet to be demystified. MAP7 made this quote resonate with me like never before: "The more you know, the more you realize you don't know." by Aristotle.
Thank you so much for reading!
Lots of Love,
Lisa
P.S. I would like to dedicate this post to Jaime Fraser and Luke Ferro, the best collaborators I have ever had, by far. Also, the beautiful banner art is by Janet Iwasa, please check out her lab page for more beautiful designs.
Please reference my thesis for more information!